Meet Loop assembly
Standardised Type IIS and overlap assembly, combined.
Pollak, B. , Cerda, A. , Delmans, M. , Álamos, S. , Moyano, T. , West, A. , Gutiérrez, R. A., Patron, N. , Federici, F. and Haseloff, J. (2019), Loop assembly: a simple and open system for recursive fabrication of DNA circuits. New Phytologist 222. doi:10.1111/nph.15625
Combinatorial and exponential assembly
Place genetic modules, sort them as you like and compose sophisticated genetic constructs in less than a week.
Compact plasmid library
An extremely compact library of 8 plasmids in two different levels of assembly yields tractable and reliable plasmid stocks.
Streamlined recursive logic
A streamlined logic using standard overhangs enable recursive assembly by looping between Odd and Even levels of assembly.
Standardised overlap assembly
Standardised Unique Nucleotide Sequences (UNSs) enable standardised overlap assembly (such as Gibson assembly) with reusable primer pairs.
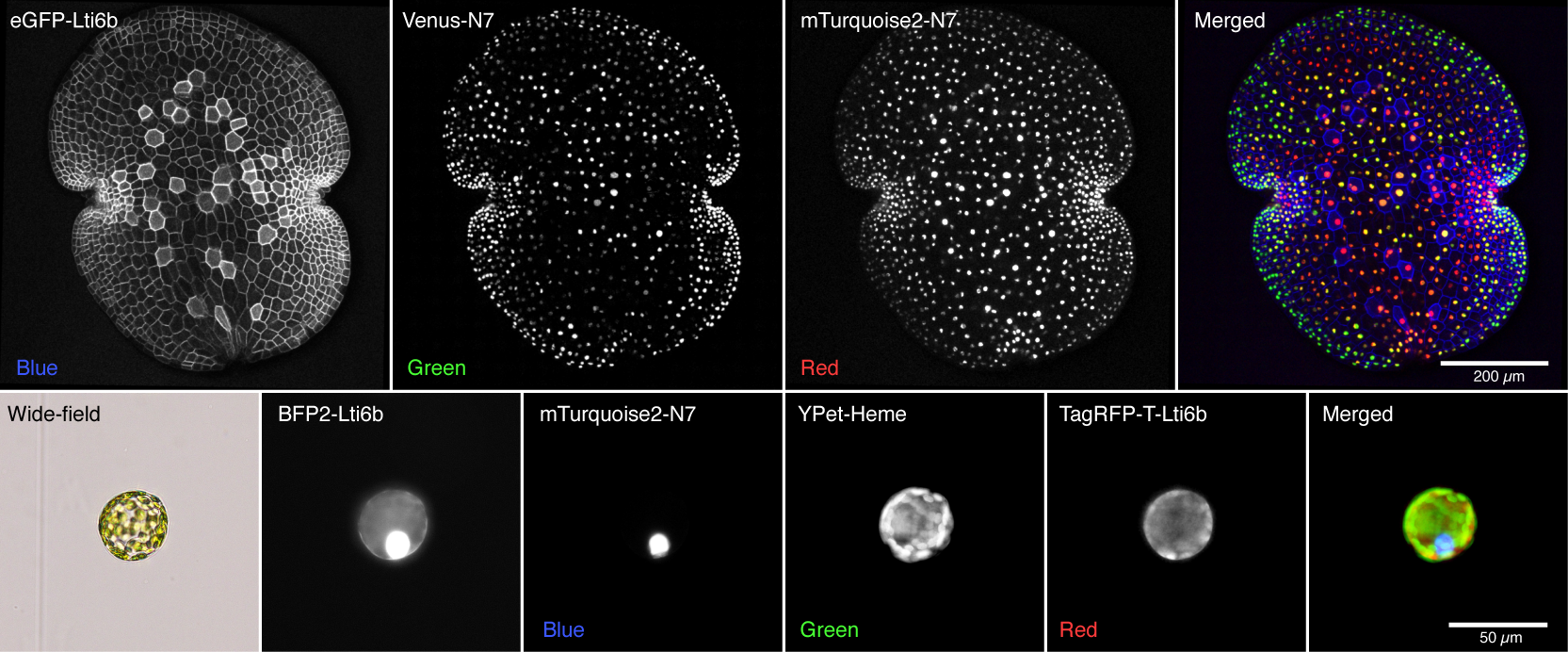
(Upper panel) A Marchantia polymorpha gemma stable transformant expressing three fluorescent reporters by Bernardo Pollak.
(Lower panel) An Arabidopsis thaliana showing transient expression of four fluorescent reporters by Ariel Cerda.
Multi-gene constructs
Obtain stable expression of multiple heterologous genes in planta.




